Remote files and fsspec caching¶
Author: Luis Lopez @betolink
When working with large binary files stored in the cloud, tools like fsspec play a critical role in abstracting file access. These tools are input/output drivers to file systems. Libraries that understand how to read these binary formats do not know anything about remote files and fsspec is in charge of translating byte requests to remote byte requests (HTTP).
Most of the file formats in the scientific world were not designed for efficient cloud access as they require non-sequential reads, meaning client libraries can jump to specific byte ranges to read only what's needed, like a row group in a Parquet file or a variable slice in NetCDF. However, the way fsspec fetches data from remote sources is controlled by its caching strategy. This strategy determines how data is fetched, stored locally, and re-used. If this strategy isn't properly aligned with the access patterns of the file format, performance and reliability can (and will) quickly degrade.
Imagine reading a NetCDF file hosted on S3 with a default block cache of 5 MB. If a query requires multiple small reads scattered throughout the file, and each of these reads falls in a different 5 MB block, fsspec will repeatedly download new blocks, resulting in excessive data transfer and latency. Worse, if the file is very large and caching is disabled or limited (e.g. using simplecache with no chunk awareness), access may become painfully slow — or even fail if many tiny reads cause timeouts or bandwidth limits to be exceeded.
By tuning the cache size, choosing the right caching backend (like blockcache or filecache), and understanding how your format is read by your analysis tools (e.g., xarray, pandas, pyarrow), you can greatly reduce redundant downloads, improve throughput, and make interactive analysis over the cloud viable.
Aligning cache strategies on a per-file basis is not a simple task, therefore earthaccess won't try to fine tune your requests except for changing the default behavior in fsspec from readahead to blockcache that offers better access patterns and a more efficient cache invalidation strategy for binary files. Based on file size earthaccess will cache starting at 4MB up to 16MB for very large files (more than 1GB).
The max in-memory caching size will be 16MBx32 blocks = 512MB per file. All of this is configurable at runtime with the new open_kwargs parameter. Some examples of the usage of this new config dictionary are detailed in this notebook.
- Efficiency: Avoids repeated downloads of the same data during random-access reads.
- Performance: Reduces latency and bandwidth usage, especially for cloud-hosted files.
- Scalability: Supports parallel or repeated access patterns without overloading remote servers.
NetCDF and HDF5 files¶
A considerable amount of remote sensing data is stored in HDF5 and NetCDF formats. They are hierarchical, self-describing formats widely used in Earth science due to their ability to store multidimensional arrays, metadata, and complex data structures efficiently. However, their design assumes relatively fast random access to local storage, which becomes a bottleneck when accessed over high-latency networks like HTTP or S3.
HDF5 and NetCDF files are typically organized into chunks—contiguous blocks of data that are compressed and stored separately. When reading a subset of a dataset (e.g., a time slice or spatial region), only the relevant chunks should be fetched. But without proper caching, each byte-range request for a chunk can result in a separate HTTP GET request, leading to thousands of small requests (Ro to Rn in the figure) for large analyses. The diagram below illustrates how traditional HDF5 access results in scattered, inefficient requests, while cloud-optimized layouts enable sequential or sparse access with fewer round trips:
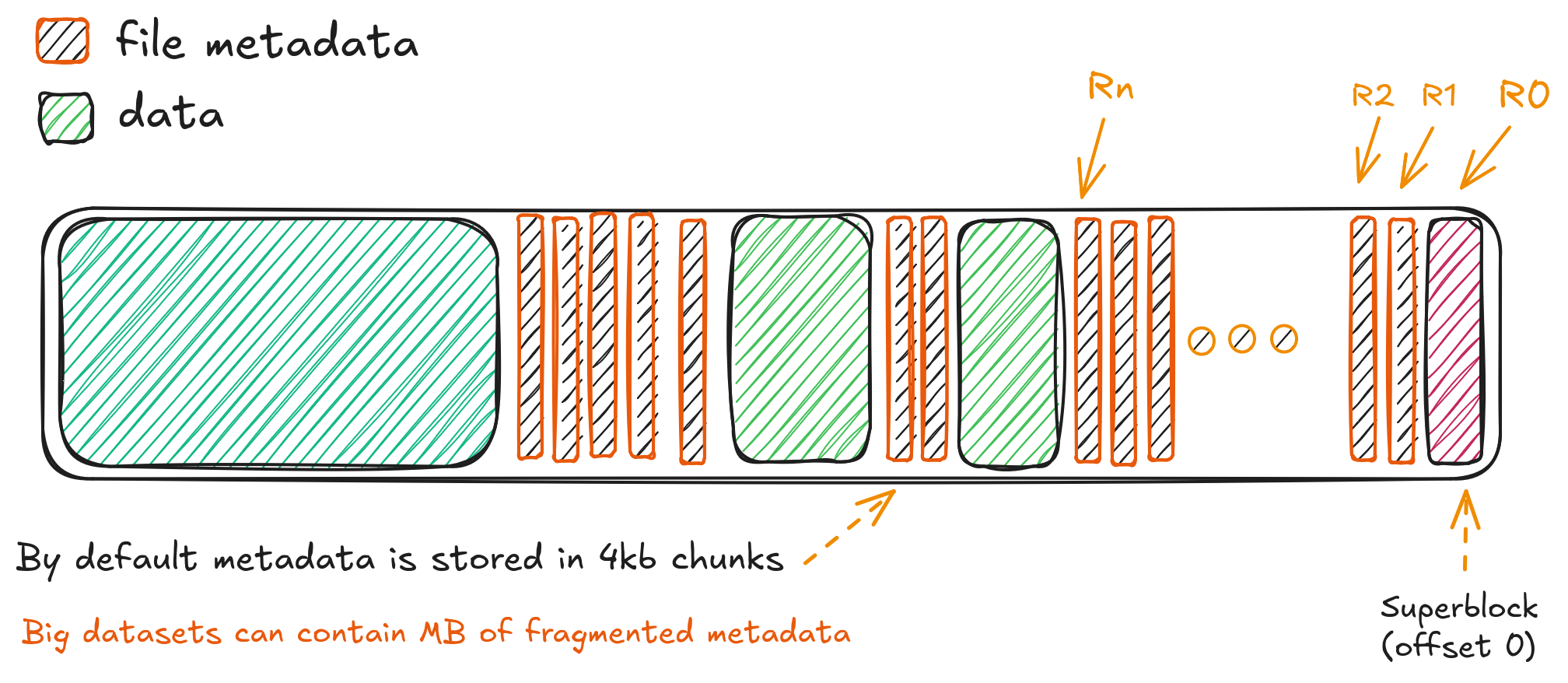
This is where fsspec's blockcache becomes essential. By aligning the cache block size with the HDF5/NetCDF chunk size (or a multiple thereof), we minimize redundant fetches and ensure that once a chunk is downloaded, it remains available for reuse during subsequent accesses. For example, if a NetCDF variable uses 1 MB chunks, a 4–16 MB block cache ensures that multiple chunks are retained per block, reducing round trips.
Additionally, because HDF5 files often contain shared metadata (e.g., dimensions, attributes, group structures), caching the beginning of the file (where some of this metadata resides) can dramatically speed up repeated opens or inspections. earthaccess leverages blockcache with sensible defaults to preserve both data and metadata across requests, making workflows using xarray or h5py over S3 much more performant.
Cloud optimized files (Zarr / COG / etc / Co-HDF5)¶
As cloud-native workflows mature, new formats and adaptations have emerged to address the limitations of traditional binary formats in object storage environments. These cloud-optimized formats are designed for efficient, parallel, and distributed access over HTTP/S3 with minimal latency.
Zarr: A chunked, compressed, and directory-structured format that natively supports cloud storage. Each chunk is stored as a separate object, enabling true parallel I/O and eliminating the need for byte-range requests. When used with fsspec, Zarr datasets can be read efficiently even with minimal caching, since each chunk is an independent resource.
Cloud Optimized GeoTIFF (COG): A GeoTIFF variant with internal tiling and overviews organized so that key image metadata is at the front of the file, and tiles are laid out sequentially. This allows fast previews and spatial subsetting via byte-range requests without downloading the entire file.
Co-HDF5 (Cloud-Optimized HDF5): An emerging pattern where HDF5 files are rechunked and reorganized to improve cloud access performance. This includes placing metadata at the beginning, aligning chunks to network block boundaries, and minimizing cross-chunk references.
These formats work best when paired with intelligent caching strategies. While Zarr benefits less from large block caches (since each chunk is already a discrete object), COG and Co-HDF5 still rely heavily on byte-range requests and thus benefit significantly from blockcache with appropriately sized blocks.
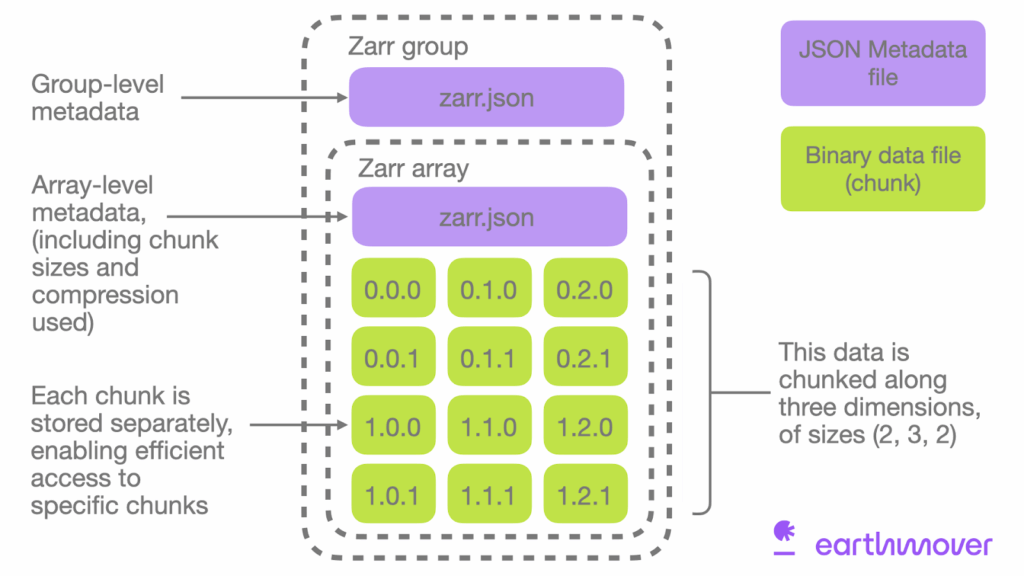
Credit: Earthmover "What is Zarr"
Ultimately, the key to high-performance cloud data access lies in matching the caching strategy to the data layout, access pattern, and underlying format. With earthaccess, we aim to provide defaults that "just work" while giving advanced users full control via open_kwargs to tune behavior as needed.
💡 Pro Tip: Always inspect your data’s chunking structure (e.g., using
zarr.tree()orh5py.File(...).visititems()) before setting cache parameters. Aligning cache and chunk sizes can yield 10x+ performance improvements.
xarray and earthaccess.open()¶
xarray has become the de facto library to open and analyze n-dimensional scientific data in Python, thus the use of fsspec to open our remote files has the advantage of being interoperable with any xarray backend(or file engine) that can accept "file-like" objects.
import earthaccess as ea
import xarray as xr
ea.login()
<earthaccess.auth.Auth at 0x7b4706c50ad0>
results = ea.search_data(
short_name="ATL03",
version="006",
cloud_hosted=True,
temporal=("2025-01", "2025-08"),
bounding_box=(33.98, 31.19, 34.65, 31.51),
count=1,
)
results[0]
Now that we have results from CMR we can open the files, in this case a single HDF5 level 2 (non gridded) swath.
One thing that this particular collection will highlight is that when dealing with non cloud-optimized files each dimension and variable will add overhead on the range requests. ATL03 contains more than 900 of them, and if we use libraries like xarray that try eagerly to fetch metadata to decode variables and dimensions our open_dataset() operation in xarray will be exponentially slow.
Let's open the file and inspect the new caching defaults from earthaccess!
file_handler = ea.open(results)[0] # only one item in the list
file_handler.f.cache
<BlockCache:
block size : 8388608
block count : 65
file size : 542254177
cache hits : 0
cache misses: 0
total requested bytes: 0>
Here we are seeing the new caching defaults when we open files using earthaccess, instead of readahead we are using the blockcache and earthaccess adjusted the block size to 8MB. This will helps us not re-fetching data when we in turn use xr.open_dataset()
%%time
# no engine defined, xarray will guess based on file type in this case h5netcdf
ds = xr.open_dataset(file_handler, group="/gt1l/heights")
ds
CPU times: user 545 ms, sys: 178 ms, total: 723 ms Wall time: 33.3 s
<xarray.Dataset> Size: 280MB
Dimensions: (delta_time: 5595411, ds_surf_type: 5)
Coordinates:
* delta_time (delta_time) datetime64[ns] 45MB 2025-01-04T00:36:08.1495...
lat_ph (delta_time) float64 45MB ...
lon_ph (delta_time) float64 45MB ...
Dimensions without coordinates: ds_surf_type
Data variables:
dist_ph_across (delta_time) float32 22MB ...
dist_ph_along (delta_time) float32 22MB ...
h_ph (delta_time) float32 22MB ...
pce_mframe_cnt (delta_time) uint32 22MB ...
ph_id_channel (delta_time) uint8 6MB ...
ph_id_count (delta_time) uint8 6MB ...
ph_id_pulse (delta_time) uint8 6MB ...
quality_ph (delta_time) int8 6MB ...
signal_conf_ph (delta_time, ds_surf_type) int8 28MB ...
weight_ph (delta_time) uint8 6MB ...
Attributes:
Description: Contains arrays of the parameters for each received photon.
data_rate: Data are stored at the photon detection rate.Better caching¶
xarray decoded a few variables here and touched many parts of the file, what does this mean regardless file access? if we inspect the cache again will see the number of requests made and how many of them hit a cached block.
file_handler.f.cache
<BlockCache:
block size : 8388608
block count : 65
file size : 542254177
cache hits : 1762
cache misses: 22
total requested bytes: 184549376>
Defaults are bad for random access: testing readahead on HDF5, ATL03 data¶
Now let's explore what happens with fsspec defaults by overriding the config with open_kwargs.
default_fsspec_open = {
"cache_type": "readahead",
"block_size": 5 * 1024 * 1024, # 5MB
}
file_handler = ea.open(results, open_kwargs=default_fsspec_open)[0]
file_handler.f.cache
<ReadAheadCache:
block size : 5242880
block count : 0
file size : 542254177
cache hits : 0
cache misses: 0
total requested bytes: 0>
%%time
ds = xr.open_dataset(file_handler, group="/gt1l/heights")
# we print the cache info again
file_handler.f.cache
CPU times: user 6.06 s, sys: 1.54 s, total: 7.6 s Wall time: 11min 2s
<ReadAheadCache:
block size : 5242880
block count : 0
file size : 542254177
cache hits : 1158
cache misses: 604
total requested bytes: 3120435184>
This we can see we requested an order of magnitude more information with considerable more cache misses despite the block_size being similar in size. Most of it is due how readahead does cache invalidation for random access.
Old defaults:
- 3120435184 requested bytes
- 604 cache misses
- Wall time: 11min
New defaults:
- 184549376 requested bytes
- 22 cache misses
- Wall time: 29.7 s
print(f"{int(3120435184 / 184549376)} times more data requested.")
16 times more data requested.
Here we are seeing over an order of magnitude improvement without changing anything but the way we fetch data. Of course things can improve even further if we deal with netCDF4/HDF5 files with fewer variables and/or especially with cloud-optimized HDF5.
We can test that with version 7 of the same file.
results = ea.search_data(
short_name="ATL03",
version="007",
cloud_hosted=True,
temporal=("2025-01", "2025-08"),
bounding_box=(33.98, 31.19, 34.65, 31.51),
count=1,
)
results[0]
file_handler = ea.open(results, open_kwargs=default_fsspec_open)[0]
file_handler.f.cache
<ReadAheadCache:
block size : 5242880
block count : 0
file size : 570425344
cache hits : 0
cache misses: 0
total requested bytes: 0>
%%time
ds = xr.open_dataset(file_handler, group="/gt1l/heights")
# we print the cache info again
file_handler.f.cache
CPU times: user 316 ms, sys: 55.7 ms, total: 371 ms Wall time: 16.2 s
<ReadAheadCache:
block size : 5242880
block count : 0
file size : 570425344
cache hits : 1274
cache misses: 13
total requested bytes: 68816555>
Wait, what happened now?¶
We didn't optimize the cache and yet this file in version 7 took only ~17 seconds to open, why? well this is a cloud optimized HDF5 file, hence all the metadata fits in the first request fsspec does. But this is not always the case, depending on how big a file is and how many variables and metadata has. Now we's use new earthaccess defaults and see if we can improve this even more.
Note that cloud optimized HDF5 and NetCDF4 is a very recent and for now not a lot of datasets will be released in this format
%%time
# best case scenario, block caching on a cloud optimized HDF5 file.
file_handler = ea.open(results)[0]
ds = xr.open_dataset(file_handler, group="/gt1l/heights")
file_handler.f.cache
CPU times: user 217 ms, sys: 52.3 ms, total: 269 ms Wall time: 6.71 s
<BlockCache:
block size : 8388608
block count : 68
file size : 570425344
cache hits : 1287
cache misses: 3
total requested bytes: 25165824>
Benchmarks¶
blockcacheis better thanreadaheadfor binary files (HDF5, netCDF4, HDF) that require random access. earthaccess now usesblockcacheby default and the block size is adjusted by file size.- small files < 100MB: 4MB
- medium files < 1GB: 8MB
- big file: > 1GB: 16MB
- optimal caching depends very much on the access pattern and the internal chunking of a given dataset, as of now that information is not easily available beforehand, a database that contains this kind of information will be really helpful to the community.
Here is a table that shows the performance gains by this new caching schema for a few selected popular binary datasets in the NASA Earth archives.
CAUTION: Running this can take up to 1 hour if we are not in-region.
%%time
import time
import earthaccess as ea
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import tqdm
import xarray as xr
ea.login()
# Some popular NASA datasets
popular_collections = [
"C2723758340-GES_DISC",
"C2724036633-LARC_CLOUD",
"C3385049989-OB_CLOUD",
"C2532426483-ORNL_CLOUD",
"C2670138092-NSIDC_CPRD",
]
collection_labels = {
"C2723758340-GES_DISC": "GPM Precipitation (GES DISC)",
"C2724036633-LARC_CLOUD": "TEMPO L2 (LARC)",
"C3385049989-OB_CLOUD": "PACE OCI (OB)",
"C2532426483-ORNL_CLOUD": "Daymet daily (ORNL)",
"C2670138092-NSIDC_CPRD": "ATL06 Lidar (NSIDC)",
}
def open_file(concept_id: str, optimized: bool, run_id: int = 0):
default_fsspec_open = {
"cache_type": "readahead",
"block_size": 5 * 1024 * 1024, # 5MB
}
results = ea.search_data(concept_id=concept_id, cloud_hosted=True, count=1)
if not results:
return {
"concept_id": concept_id,
"optimized": optimized,
"run_id": run_id,
"time": None,
"block_size": None,
"file_size": None,
"cache_hits": None,
"cache_misses": None,
"total_requested_bytes": None,
"error": "No files found",
}
try:
if not optimized:
f_h = ea.open(results, open_kwargs=default_fsspec_open)[0]
else:
f_h = ea.open(results)[
0
] # Uses earthaccess's blockcache + adaptive block size
t_start = time.time()
ds = xr.open_datatree(f_h, phony_dims="sort", decode_timedelta=True)
execution_time = time.time() - t_start
ds.close()
cache = f_h.f.cache
return {
"concept_id": concept_id,
"optimized": optimized,
"run_id": run_id,
"time": execution_time,
"block_size": getattr(cache, "blocksize", None),
"file_size": getattr(cache, "size", None),
"cache_hits": getattr(cache, "hit_count", 0),
"cache_misses": getattr(cache, "miss_count", 0),
"total_requested_bytes": getattr(cache, "total_requested_bytes", 0),
"error": None,
}
except Exception as e:
return {
"concept_id": concept_id,
"optimized": optimized,
"run_id": run_id,
"time": None,
"block_size": None,
"file_size": None,
"cache_hits": None,
"cache_misses": None,
"total_requested_bytes": None,
"error": str(e),
}
n_runs = 3
print(f"Starting benchmark: {n_runs} runs per dataset and mode\n")
benchmarks = []
for run in range(n_runs):
print(f"Run {run + 1} of {n_runs}")
# Optimized (earthaccess defaults)
print("Optimized (blockcache)")
for cid in tqdm.tqdm(popular_collections):
result = open_file(cid, optimized=True, run_id=run)
benchmarks.append(result)
# Default (readahead)
print("Non-optimized (readahead)")
for cid in tqdm.tqdm(popular_collections):
result = open_file(cid, optimized=False, run_id=run)
benchmarks.append(result)
df = pd.DataFrame(benchmarks)
Starting benchmark: 3 runs per dataset and mode Run 1 of 3 Optimized (blockcache)
100%|███████████████████████████████████████████████████████████████████████| 5/5 [01:42<00:00, 20.48s/it]
Non-optimized (readahead)
100%|██████████████████████████████████████████████████████████████████████| 5/5 [11:46<00:00, 141.24s/it]
Run 2 of 3 Optimized (blockcache)
100%|███████████████████████████████████████████████████████████████████████| 5/5 [01:49<00:00, 21.98s/it]
Non-optimized (readahead)
100%|██████████████████████████████████████████████████████████████████████| 5/5 [12:01<00:00, 144.28s/it]
Run 3 of 3 Optimized (blockcache)
100%|███████████████████████████████████████████████████████████████████████| 5/5 [01:36<00:00, 19.24s/it]
Non-optimized (readahead)
100%|██████████████████████████████████████████████████████████████████████| 5/5 [12:21<00:00, 148.27s/it]
CPU times: user 54.4 s, sys: 3.94 s, total: 58.3 s Wall time: 41min 18s
df["dataset"] = df["concept_id"].map(collection_labels)
df["mode"] = df["optimized"].map({True: "Optimized", False: "Default (readahead)"})
df["file_size_mb"] = df["file_size"] / (1024 * 1024)
df["total_requests"] = df["cache_hits"] + df["cache_misses"]
df["hit_rate"] = df["cache_hits"] / df["total_requests"].where(
df["total_requests"] > 0, 1
)
df["requested_mb"] = df["total_requested_bytes"] / (1024 * 1024)
success_df = df[df["error"].isna()].copy()
failed = df[df["error"].notna()]
if len(failed) > 0:
print(failed[["dataset", "mode", "run_id", "error"]].drop_duplicates())
if len(success_df) == 0:
raise RuntimeError("All tests failed. Check authentication and connectivity.")
# Average metrics over n runs
grouped = (
success_df.groupby(["dataset", "mode"])
.agg(
time_mean=("time", "mean"),
time_std=("time", "std"),
hit_rate_mean=("hit_rate", "mean"),
requested_mb_mean=("requested_mb", "mean"),
block_size=("block_size", "first"), # should be stable
file_size_mb=("file_size_mb", "first"),
)
.reset_index()
)
pivot = grouped.pivot(index="dataset", columns="mode")
pivot.columns = [f"{metric}_{mode}" for metric, mode in pivot.columns]
pivot = pivot.sort_index()
pivot["speedup_x"] = (
pivot["time_mean_Default (readahead)"] / pivot["time_mean_Optimized"]
).round(2)
pivot["data_savings_%"] = (
(
pivot["requested_mb_mean_Default (readahead)"]
- pivot["requested_mb_mean_Optimized"]
)
/ pivot["requested_mb_mean_Default (readahead)"]
* 100
).round(1)
final_cols = [
"time_mean_Optimized",
"time_mean_Default (readahead)",
"speedup_x",
"hit_rate_mean_Optimized",
"hit_rate_mean_Default (readahead)",
"requested_mb_mean_Optimized",
"requested_mb_mean_Default (readahead)",
"data_savings_%",
"block_size_Optimized",
"block_size_Default (readahead)",
"file_size_mb_Optimized",
]
summary = pivot[final_cols].copy()
summary.rename(
columns={
"time_mean_Optimized": "Time (s) - Opt",
"time_mean_Default (readahead)": "Time (s) - Default",
"speedup_x": "Speedup (x)",
"hit_rate_mean_Optimized": "Hit Rate - Opt",
"hit_rate_mean_Default (readahead)": "Hit Rate - Default",
"requested_mb_mean_Optimized": "Req MB - Opt",
"requested_mb_mean_Default (readahead)": "Req MB - Default",
"data_savings_%": "Data Saved (%)",
"block_size_Optimized": "Block MB - Opt",
"block_size_Default (readahead)": "Block MB - Default",
"file_size_mb_Optimized": "File Size (MB)",
},
inplace=True,
)
for col in ["Block MB - Opt", "Block MB - Default"]:
summary[col] = (summary[col] / (1024 * 1024)).round(1)
# Summary
print("\n" + "=" * 100)
print(f"SUMMARY: Averaged Over {n_runs} Runs")
print("=" * 100)
print(summary.round(3))
# Cache Hit Rate Comparison
plot_data = grouped.set_index(["dataset", "mode"])["hit_rate_mean"].unstack("mode")
datasets = plot_data.index
x = np.arange(len(datasets))
width = 0.35
plt.figure(figsize=(10, 6))
bars1 = plt.bar(
x - width / 2,
plot_data["Default (readahead)"],
width,
label="Default (readahead)",
color="gray",
alpha=0.8,
)
bars2 = plt.bar(
x + width / 2,
plot_data["Optimized"],
width,
label="Optimized (blockcache)",
color="seagreen",
alpha=0.9,
)
plt.ylabel("Cache Hit Rate", fontsize=12)
plt.title(
"Cache Hit Rate: Optimized vs Default (Averaged Over 3 Runs)", fontsize=14, pad=20
)
plt.xticks(x, [lbl.replace(" ", "\n") for lbl in datasets], rotation=0)
plt.ylim(0, 1.05)
plt.legend(fontsize=11)
plt.grid(axis="y", linestyle="--", alpha=0.4)
def add_labels(bars):
for bar in bars:
height = bar.get_height()
plt.text(
bar.get_x() + bar.get_width() / 2,
height + 0.02,
f"{height:.3f}",
ha="center",
va="bottom",
fontsize=9,
)
add_labels(bars1)
add_labels(bars2)
plt.tight_layout()
plt.show()
====================================================================================================
SUMMARY: Averaged Over 3 Runs
====================================================================================================
Time (s) - Opt Time (s) - Default Speedup (x) \
dataset
ATL06 Lidar (NSIDC) 10.357 507.886 49.04
Daymet daily (ORNL) 2.406 2.025 0.84
GPM Precipitation (GES DISC) 4.206 57.330 13.63
PACE OCI (OB) 3.581 24.653 6.88
TEMPO L2 (LARC) 19.316 68.545 3.55
Hit Rate - Opt Hit Rate - Default \
dataset
ATL06 Lidar (NSIDC) 0.999 0.810
Daymet daily (ORNL) 0.998 0.998
GPM Precipitation (GES DISC) 0.979 0.452
PACE OCI (OB) 0.990 0.887
TEMPO L2 (LARC) 0.916 0.592
Req MB - Opt Req MB - Default Data Saved (%) \
dataset
ATL06 Lidar (NSIDC) 12.0 1996.835 99.4
Daymet daily (ORNL) 4.0 1.922 -108.1
GPM Precipitation (GES DISC) 8.0 134.247 94.0
PACE OCI (OB) 8.0 67.849 88.2
TEMPO L2 (LARC) 112.0 288.875 61.2
Block MB - Opt Block MB - Default \
dataset
ATL06 Lidar (NSIDC) 4.0 5.0
Daymet daily (ORNL) 4.0 5.0
GPM Precipitation (GES DISC) 4.0 5.0
PACE OCI (OB) 4.0 5.0
TEMPO L2 (LARC) 8.0 5.0
File Size (MB)
dataset
ATL06 Lidar (NSIDC) 11.307
Daymet daily (ORNL) 1.922
GPM Precipitation (GES DISC) 7.276
PACE OCI (OB) 8.435
TEMPO L2 (LARC) 292.735
TL;DR; earthaccess.open() then vs now¶
All of this to say that with the upcoming release of earthaccess we'll enjoy speedups in our workflows from marginal to order of magnitude depending on the dataset.
plt.figure(figsize=(10, 6))
# Extract data
speedup_values = summary["Speedup (x)"].values
datasets = summary.index
# Color: red if < 1.0 (slower), green if >= 1.0 (faster)
colors = ["firebrick" if s < 1.0 else "seagreen" for s in speedup_values]
bars = plt.bar(
datasets, speedup_values, color=colors, alpha=0.85, edgecolor="black", linewidth=0.8
)
# Labels and title
plt.ylabel("Speedup Factor (x)", fontsize=12)
plt.title("Speedup: Optimized vs Default (Higher = Better)", fontsize=14, pad=20)
plt.xticks(rotation=0)
plt.grid(axis="y", linestyle="--", alpha=0.5)
plt.axhline(1.0, color="red", linestyle="--", linewidth=1.2, label="No difference")
# Add value labels on bars
for bar, speedup in zip(bars, speedup_values):
color = "white" if speedup < 1.0 else "black"
plt.text(
bar.get_x() + bar.get_width() / 2,
bar.get_height() + max(speedup_values) * 0.05,
f"{speedup:.2f}x",
ha="center",
va="bottom",
fontsize=10,
color=color,
fontweight="bold",
)
plt.legend(fontsize=11)
plt.tight_layout()
plt.show()



